rna hydrolysis rate
|
Measurement of the rate of RNA hydrolysis in aqueous solution at
We have designed an unique reaction system to analyze hydrolysis rate of phosphodiester linkage at elevated temperatures using oligonucleotides with sequence of |
|
Non-Enzymatic RNA Hydrolysis Promoted by the Combined
RNA RNA Degradation |
|
RNA hydrolysis by cobalt(III) complexes1
probe to assign the rate-limiting step in RNA hydrolysis. A novel mechanism involving general acid catalysis by CoIII- bound water is proposed on the basis |
|
Central role of the RNA polymerase trigger loop in intrinsic RNA
15 Jun 2010 The active center of RNA polymerase can hydrolyze phospho- ... effects on the rate of second phosphodiester bond hydrolysis. |
|
Duplex Structure of Double-Stranded RNA Provides Stability against
25 May 2021 Comparing the rate constant for. dsRNA hydrolysis to that for ssRNA hydrolysis indicates that. dsRNA was more resistant to alkaline hydrolysis ... |
|
Reconsidering the energetics of ribonuclease catalysed RNA
From the uncatalysed rates of UpA cleavage [23] and dimethyl phosphate (as a model for DNA) hydrolysis [24] we estimate the rate at which RNA depolymerizes in |
|
Theoretical basis for stabilizing messenger RNA through secondary
24 Aug 2020 reduce mRNA hydrolysis is to redesign RNAs to form double-stranded regions ... model that links an RNA molecule's overall hydrolysis rate to ... |
|
RNA hydrolysis by cobalt(III) complexes1
probe to assign the rate-limiting step in RNA hydrolysis. A novel mechanism involving general acid catalysis by CoIII- bound water is proposed on the basis |
|
Excess dNTPs Minimize RNA Hydrolysis during Reverse Transcription
buffers (3) the rate of RNA breakdown rate of loss of intact 7.5-kb RNA was determined |
|
THE KINETICS OF THE ENZYMATIC HYDROLYSIS OF
Numerous methods have been utilized to follow the rate of the enzy- matic hydrolysis of RNA.' Bain and Rusch (1) measured the liberation. |
|
Role of the RNA polymerase trigger loop in catalysis and pausing
Figure 2The TL is dispensable for intrinsic RNA hydrolysis on a locked scaffold (a) Nucleic-acid scaffold DNA and RNA are shown in black and red respectively Red asterisk 32P label (b) |
|
Phosphodiester bond - Wikipedia
RNAhydrolysis was detectable with 3 91 nM RNase A (Fig 3c)indicating a sensitivity of *5 ng (100ll of 3 91 nMRNase A corresponds to 5 4 ng of protein) This methodseems to be*400-fold more sensitive than the traditionalin situ gel electrophoresis-based method [13] whichrequires*2lg of protein |
|
RNA Folding and Hydrolysis Terms Explain ATP - Manoharan
human systems By examining the basic thermodynamic equations for RNA interactions in RNAi we demonstrate how the free energies of RNA folding and phosphoester bond hydrolysis can drive RNAi without ATP Our calculations of RNAi efficiency are close to actual values obtained from in vitro experimental data from two previous studies for |
|
Searches related to rna hydrolysis rate filetype:pdf
To determine whether the rate of RNA binding limited the observed rate of hydrolysis of the RNA substrate we performed a rapid quench experiment with the dsRNA substrate using two |
What is the relative ease of RNA hydrolysis?
- The relative ease of RNA hydrolysis is an effect of the presence of the 2' hydroxyl group . A phosphodiesterase is an enzyme that catalyzes the hydrolysis of phosphodiester bonds, for instance a bond in a molecule of cyclic AMP or cyclic GMP . An enzyme that plays an important role in the repair of oxidative DNA damage is the 3'-phosphodiesterase.
What is the rate of hydrolysis?
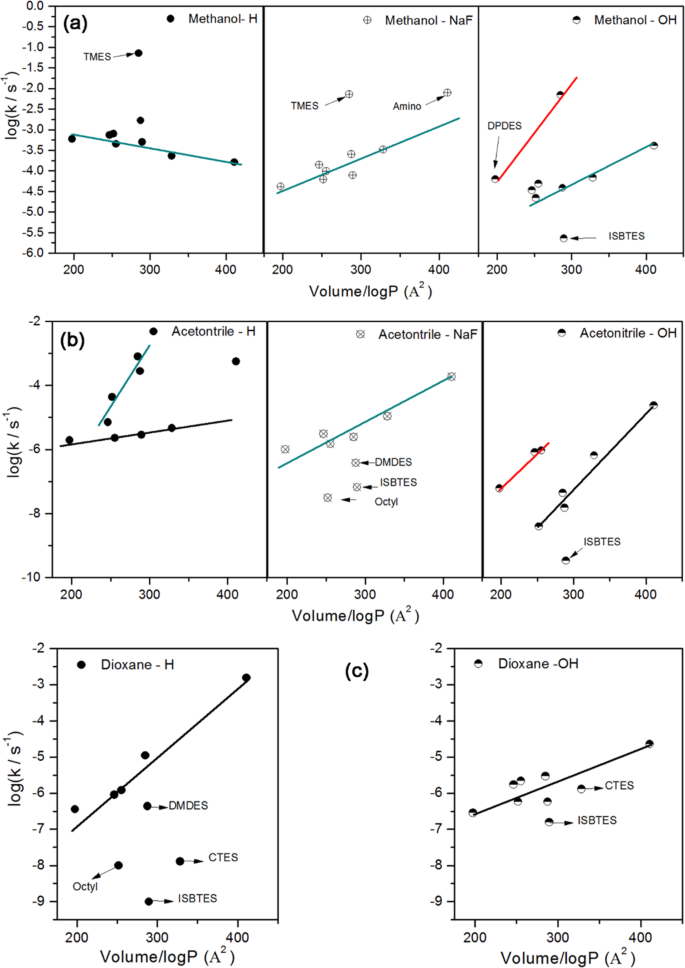
- The rate of hydrolysis depends on the concentration of silane, the pH of the solution (pH > 10 and pH < 5 give a high rate, pH near 7 gives the slowest rate), and the character of the functional groups of silanes and alkoxy groups [ 20 – 22 ]. Hydrolysis is catalyzed by hydronium ions H 3 O +, hydroxyl ions OH ?, Lewis bases, and some metals.
Why is RNA not susceptible to base-catalyzed hydrolysis?
- RNA is susceptible to this base-catalyzed hydrolysis because the ribose sugar in RNA has a hydroxyl group at the 2’ position. This feature makes RNA chemically unstable compared to DNA, which does not have this 2’ -OH group and thus is not susceptible to base-catalyzed hydrolysis. Mechanism of base catalyzed RNA hydrolysis.
Where does RNA hydrolyze in an alkaline environment?
- The hydrolysis occurs at the phosphodiester bond of the backbone. Hence, RNA rapidly hydrolyzes in an alkaline environment into cyclic 2',3'-monophosphate nucleotides, which undergo further hydrolysis in water to form a mixture of 2'- and 3'-nucleoside monophosphates. This breaking of the backbone of RNA is known as a strand break.
|
Measurement of the rate of RNA hydrolysis in aqueous solution at
42 289-290 Measurement of the rate of RNA hydrolysis in aqueous solution at elevated temperatures using a new monitoring method for hydrothermal reactions |
|
Enormously Fast RNA Hydrolysis by Lanthanide(III) Ions under
The hydrolysis rate is 105-fold faster than that of ApA (the rate constants at pH 8 0 are 7 x 10~3 and 7 X 10~8 min"1, respectively) |
|
Comparing Spontaneous Hydrolysis Rates of Activated - Zenodo
Abstract—This research project aims to investigate difference in relative rates concerning phosphoryl transfer relevant to biological catalysis of DNA and RNA in |
|
THE KINETICS OF THE ENZYMATIC HYDROLYSIS - ScienceDirect
2 for the three lower enzyme concentrations Effect of Substrate Concentration- The hydrolysis of RNA was measured at a series of substrate levels, and initial rates |
|
Hydrolysis of Double-Stranded and Single-Stranded RNA in Hairpin
ReceiVed August 8, 1996 RNA is hydrolyzed in ViVo by ribonucleases via a two- step increase the rate of hydrolysis and tune the selectivity of the catalyst |
|
Theoretical basis for stabilizing messenger RNA through - bioRxiv
24 août 2020 · average unpaired probability of an mRNA, or AUP, to its overall rate of hydrolysis To characterize the stabilization achievable through structure |
|
A neutral pH thermal hydrolysis method for - RNA Journal
23 mai 2014 · Thermal hydrolysis can be performed at neutral pH without reaction quenching, the rate of cytosine deamination in DNA at 100°C is slowest |
|
Hydrolysis of DNA and RNA by lanthanide ions - Shigekawa Lab
(6) The nucleic acid bases do not directly participate in the catalysis (the rate of DNA hydrolysis is almost independent of them) (7) The reaction is accompanied by |

![Kinetics of hydrolysis of diltiazem - [PDF Document] Kinetics of hydrolysis of diltiazem - [PDF Document]](https://0.academia-photos.com/attachment_thumbnails/46461544/mini_magick20190209-8382-12fwmur.png?1549766175)






















.jpg)




