 Protocol: Hemocytometer Cell Counting
Protocol: Hemocytometer Cell Counting
19 июл. 2019 г. In general boxes 1-4 (red) are used to count cells. 1 Trypan Blue-cell suspension dilution may change based upon predicted cell concentration.
 Countess™ II FL Automated Cell Counter
Countess™ II FL Automated Cell Counter
26 окт. 2015 г. Therefore the cell concentration displayed in the Results screen must be multiplied by the dilution factor to calculate the original cell ...
 Cell Counting using a Hemocytometer Based on YouTube video
Cell Counting using a Hemocytometer Based on YouTube video
Examples of when to count cells: • Passaging: you may not need to count every time but can be helpful to determine how many flasks to seed
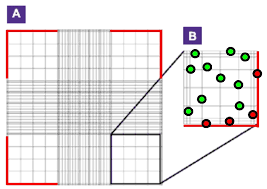 Cell count and viability assay Grid Colour Area Volume 3x3 Blue 1
Cell count and viability assay Grid Colour Area Volume 3x3 Blue 1
10 мар. 2008 г. The stained cells are immediately counted under microscopic observation using a Neubauer haemocytometer. Although the original protocol suggests ...
 Measuring absolute count with absolute certainty - Measurement
Measuring absolute count with absolute certainty - Measurement
are available for cell counting methodologies. Figure 2: (A) Measured cell count on a MACSQuant Analyzer as a function of dilution factor on a log-log scale.
 Algae to Energy - Using and Re-using a Hemocytometer to Count
Algae to Energy - Using and Re-using a Hemocytometer to Count
Hemocytometer to Count Algae Cells. 1). Prepare your sample by shaking your photobioreactor for at least 30 seconds and use a 4). Calculate cell density. ...
 Celgene PowerPoint Presentation With Confidentiality Notice
Celgene PowerPoint Presentation With Confidentiality Notice
10 апр. 2017 г. Ideal cell count. Typical cell count due to lack of cell reference standard ... • Choice of diluents and dilution factor impact cell counting.
 Automated Hemocytometer-Based Live/Dead Cell Counting using
Automated Hemocytometer-Based Live/Dead Cell Counting using
Often an exclusion dye such as trypan blue is used such that live cells can be differentiated from dead cells and the cell count reflects only viable cells.
 TECHNICAL NOTE - Guidelines for Accurate Target Cell Counts
TECHNICAL NOTE - Guidelines for Accurate Target Cell Counts
Hemocytometer Cell Count Distributions: Implications of Non-Poisson Behavior. Biotechnology Progress 1991. Page 6. 10x Genomics®
 Trypan Blue
Trypan Blue
USE OF TRYPAN BLUE STAIN AND. THE HEMOCYTOMETER TO DETERMINE TOTAL. CELL COUNTS AND VIABLE CELL NUMBER. Trypan Blue is one of several stains recommended.
 Cell Counting 1.0 Purpose Cell counting is done in order to
Cell Counting 1.0 Purpose Cell counting is done in order to
13-Jul-2012 Cell counting is done in order to determine an accurate number of viable cells ... Average # cells counted x dilution factor (e.g. 4 for 1:4 ...
 CEREBROSPINAL FLUIDS
CEREBROSPINAL FLUIDS
CELL COUNT AND CYTOMORPHOLOGY. Dr Kiran Ghodke. Fellow Hematopathology Laboratory EDTA for cell counts and morphology ... N X Dilution factor.
 Counting mononuclear cells by method of hemocytometer
Counting mononuclear cells by method of hemocytometer
cells by method of hemocytometer. Date: October 2010. Authors: Jennifer Hossler. PURPOSE: To obtain a total mononuclear cell count collected from the
 TOTAL RED BLOOD CELL (RBC) COUNT USING
TOTAL RED BLOOD CELL (RBC) COUNT USING
06-Apr-2018 PRINCIPLE OF TOTAL RBC COUNT USING HEMOCYTOMETER. Very large numbers of Red Blood Cells are present in the Blood Specimen.
 LUNA™ Automated Cell Counter User Manual
LUNA™ Automated Cell Counter User Manual
of cell count and viability will be displayed on the screen. The counting image can be For samples with Trypan Blue Stain the dilution factor.
 CEREBROSPINAL FLUID CELL COUNTS Educational commentary
CEREBROSPINAL FLUID CELL COUNTS Educational commentary
Calculate the number of cells per microliter from the raw cell count. the calculation is even simpler because the dilution factor is 1. In this case.
 VPC Viable Plate Count
VPC Viable Plate Count
The dilution factor is used to calculate the number of cells in the original cell culture. In our example an average of 50 colonies was counted on the
 Counting cells - iGEM
Counting cells - iGEM
Calculate the cell concentration (cells/mL) of your original cell suspension: c = mean number of cells in one big 4x4 square * 10 000 * dilution factor (= 1
 WBC manual count using hemocytometer
WBC manual count using hemocytometer
To accurately count WBC in Chamber. To perform reliable dilution of blood cells. To calculate the number of cells/µL
 [PDF] Protocol: Hemocytometer Cell Counting
[PDF] Protocol: Hemocytometer Cell Counting
19 juil 2019 · In general boxes 1-4 (red) are used to count cells 1 Trypan Blue-cell suspension dilution may change based upon predicted cell concentration
 [PDF] Counting cells - iGEM
[PDF] Counting cells - iGEM
Calculate the cell concentration (cells/mL) of your original cell suspension: c = mean number of cells in one big 4x4 square * 10 000 * dilution factor (= 1
 [PDF] Calculating the Concentration of Cells - Penn State Lehigh Valley
[PDF] Calculating the Concentration of Cells - Penn State Lehigh Valley
Calculation: used to calculate cells/mL using a hemocytometer [cells/mL] (Total # cells counted) (104) (dilution factor) (Total # squares counted)
 [PDF] Counting mononuclear cells by method of hemocytometer - URMC
[PDF] Counting mononuclear cells by method of hemocytometer - URMC
(If there are to many cells to count go back and adjust dilution factor) • Using a cell counter count how many live lymphocytes there are in the four large
 [PDF] SOP 31- Cell Countingpdf
[PDF] SOP 31- Cell Countingpdf
13 juil 2012 · In order to determine total # of cells use the following equation: Average # cells counted x dilution factor (e g 4 for 1:4 dilution) x
 [PDF] Cell count and viability assay Grid Colour Area Volume 3x3 Blue 1
[PDF] Cell count and viability assay Grid Colour Area Volume 3x3 Blue 1
10 mar 2008 · haemocytometer allows an exact volume of cells to be loaded into one or two of estimate cell concentration based on direct cell counts
 [PDF] Cell Counting using Hemocytometer
[PDF] Cell Counting using Hemocytometer
Cell Counting using a Hemocytometer Based on YouTube video: https://www youtube com/watch?v=pP0xERLUhyc Examples of when to count cells:
 (PDF) Hemocytometer Calculator - ResearchGate
(PDF) Hemocytometer Calculator - ResearchGate
8 fév 2018 · PDF Cell counting is normally performed manually with a hemocytometer a detailed explanation of calculation and Observe the grid of the
 [PDF] Hemocytometerpdf
[PDF] Hemocytometerpdf
When counting count only those cells on the lines of two sides of the large square to avoid counting cells twice (Figure 3G) Suspensions should be dilute
How do you calculate cell count by dilution factor?
The formula for dilution factor (or DF for short) is as follows: DF = (final volume of cells + stain)/(initial volume of cells). For example, If you mix your sample 1:1 with AO/PI, you'll need to add 20 uL AO/PI to 20 uL cells, for a total of 40 uL. So, DF = ( 40 uL)/(20uL cells) = 2.How do you calculate the dilution factor for the cell count hemocytometer?
Dilution Factor = Total Volume (Volume of sample + Volume of diluting liquid) / Volume of sample. Total viable cells/Sample = Viable Cells/ml x The original volume of fluid from which the cell sample was removed.- 0.5 part of blood is in 100 parts of fluid or, 1 part of blood is mixed in 200 parts of fluid Thus, dilution factor for RBC counting is 200. 0.5 part of blood is mixed in 10 parts of fluid So, 1 part of blood is in 20 parts of fluid Thus, dilution factor for WBC counting is 20.
VPC Viable Plate Count
Learning Objectives
The student will
Use aseptic techniques in the safe inoculation of various forms of media. Follow oral and written instructions and manage time in the lab efficiently. Apply correct terminology regarding microbiological techniques when making observations. Use serial dilution to estimate the concentration of microorganisms in a broth culture.Background/Theory
Estimating the number of bacterial cells in a sample, known as a bacterial count, is a commontask performed by microbiologists. The number of bacteria in a clinical sample serves as an indication of
the extent of an infection. Quality control of drinking water, food, medication, and even cosmetics relies
on estimates of bacterial counts to detect contamination and prevent the spread of disease. Two major
approaches are used to measure cell number. The direct methods involve counting cells, whereas the indirect methods depend on the measurement of cell presence or activity without actually counting individual cells. Both direct and indirect methods have advantages and disadvantages for specific applications. (OpenStax CNX, 2018)The viable plate count, or simply plate count, is a count of viable or live cells. It is based on the
principle that viable cells replicate and give rise to visible colonies when incubated under suitable
conditions for the specimen. (OpenStax CNX, 2018) A measured amount of a liquid culture is inoculated
onto a plate. The plate is incubated and the colonies that result are counted. The results are usually expressed as colony-forming units per milliliter (CFU/mL) rather thancells per milliliter because more than one cell may have landed on the same spot to give rise to a single
colony. Furthermore, samples of bacteria that grow in clusters or chains are difficult to disperse and a
single colony may represent several original cells. Some cells are described as viable but nonculturable
and will not form colonies on solid media. For all these reasons, the viable plate count is considered a
low estimate of the actual number of live cells. These limitations do not detract from the usefulness of
the method, which provides estimates of live bacterial numbers. (OpenStax CNX, 2018) There are two common approaches to inoculating plates for viable counts: the pour plate andthe spread plate methods. Although the final inoculation procedure differs between these two methods,
they both start with a serial dilution of the culture. (OpenStax CNX, 2018) Serial dilution is necessary
because the concentration of cells in even a slightly turbid culture is too large to produce discrete
colonies that are countable on a plate. The serial dilution of a culture is an important first step before proceeding to either the pourplate or spread plate method. The goal of the serial dilution process is to obtain plates with CFUs in the
range of 30-300, and the process usually involves several dilutions in multiples of 10 to simplifycalculation. The number of serial dilutions is chosen according to a preliminary estimate of the culture
density. Figure 1 illustrates the serial dilution method. (OpenStax CNX, 2018) A fixed volume of the original culture, 1.0 mL, is added to and thoroughly mixed with the first dilution tube solution, which contains 9.0 mL of sterile broth, the dilution blank. This step represents a dilution factor of 10, or 1:10, or 10-1 compared with the original culture. From this first dilution, the same volume, 1.0 mL, is withdrawn and mixed with a fresh tube of 9.0 mLof dilution solution. The dilution factor is now 1:100, or 10-2 compared with the original culture. This
VPCprocess continues until a series of dilutions is produced that will bracket the desired cell concentration
for accurate counting. (OpenStax CNX, 2018) From each tube, a sample (usually either 0.1 mL or 1.0 mL) is plated on solid medium using either the pour plate method or the spread plate method (figure 2 below). The plates are incubated until colonies appear. Thorough mixing of samples with the dilution medium (to ensure the cell distribution in the tube is random) is paramount to obtaining reliable results. (OpenStax CNX, 2018) The dilution factor is used to calculate the number of cells in the original cell culture. In our example, an average of 50 colonies was counted on the plates obtained from the 1:10,000 dilution.Because only 0.1 mL of suspension was pipetted on the plate, the multiplier (final dilution factor or FDF)
required to reconstitute the original concentration is 10 × 10,000 (final dilution factor 1:100,000 or 10-5).
The number of CFU per mL is equal to 50 × 100 × 10,000 = 5,000,000. The number of bacteria in the
culture is estimated as 5 million cells/mL. (OpenStax CNX, 2018)Another way to look at it is using this equation:
௩௨ ௧ௗ or ܦܥܱ This gives you a result with the unit CFU/mL. The volume plated can also be expressed as the final dilution factor on the plate. In figure 1, the colony count obtained from the 1:1000 dilution was 389, well below theexpected 500 for a 10-fold difference in dilutions. This highlights the issue of inaccuracy when colony
counts are greater than 300 and more than one bacterial cell grows into a single colony. (OpenStax CNX,
2018) In other words, with too much growth, colonies begin to become confluent and individual CFUs
are not represented accurately as discrete colonies. Microbiologists typically count plates with 30-300 colonies. Samples with too few colonies (<30) do not give statistically reliable numbers because of sampling error, and overcrowded plates (>300colonies) make it difficult to accurately count individual colonies. Furthermore, counts in this range
minimize occurrences of more than one bacterial cell forming a single colony. Thus, the calculated CFU is
Figure 1 Serial dilution involves diluting a fixed volume of cells mixed with dilution solution using the previous dilution as an
inoculum. The result is dilution of the original culture by an exponentially growing factor. (credit: modification of work by
͞Leberechtc"ͬWikimedia Commons) (OpenStax CNX, 2018) VPCcloser to the true number of live bacteria in the population. (OpenStax CNX, 2018) We refer to plates
with colony counts in this range as countable plates.Spread Plate Procedure
In this lab, you will plate the samples using the spread plate technique.1. Aseptically remove the cap from the dilution tube. Because this requires extra dexterity,
your partner may hold the cap. Flame the mouth of the tube as usual.2. With a sterile pipet, draw up 0.1 mL into the pipette, reflame the test tube mouth and
replace the cap. Set the tube in a rack.3. With the plate right-side-up, lift the lid enough to allow the pipet to dispense the liquid on
to the center of the plate. Discard the pipet in the disposal container.4. Remove the bent glass rod or triangular spreader from the jar of EtOH and replace the jar
lid. Pass the rod briefly through the burner flame to ignite the alcohol. Hold the horizontal length of the rod below your hand so that the burning alcohol does not run down and burn your fingers. Do not hold the rod over paper or other ignitable materials. Do not shake the rod. The alcohol will sterilize the rod. The flame merely burns off the excess.5. Spread the sample over the surface of the agar with a back and forth motion while turning
the plate. Your non-dominant hand will hold the plate lid above the plate and rotate the base while your dominant hand will use the bent rod.6. Immediately after spreading the inoculum, place the glass rod back into the alcohol jar and
replace the plate's lid. Do not set it on the bench.7. The plate should remain right-side-up for a few minutes to allow for the inoculum to soak
into the agar. It should be incubated up-side down.Experiment/Exercise
Materials per student pair
Test tube rack
5 sterile tubes
6 1.0 mL pipettes
1 10.0 mL pipette
Small pipette pump (blue)
Large pipette pump (green)
Figure 2. In the spread plate method of cell counting, the sample is poured onto solid agar and then spread using
a sterile spreader. This process is repeated for each serial dilution prepared. The resulting colonies are counted
and provide an estimate of the number of cells in the original volume samples. (OpenStax CNX, 2018) VPC4 TSA plates
1 Jar EtOH with lid
1 bent glass rod or metal spreader
Container of sterile saline
Cultures
Fresh overnight broth
E. coli
Procedure Lab 1
1. The protocol. The procedure will follow the diagram in figure 3. Fill in the tube dilution factors
and the final dilution factors on the plates in the diagram. There will be a prelab question about this diagram!Figure 3 Dilution scheme for this exercise.
2. Preparation of materials.
a. Label four plates with the DF indicated in the scheme in figure 3. To each plate, add the information required on every label in this course. b. Label five sterile tubes with the DF indicated in the scheme in figure 3 and place them in order in the rack. This is the only labeling required because these tubes will not be incubated. c. Into the 10-2 and the 10-4 sterile tubes, aseptically pipet exactly 9.9 mL sterile saline using a10.0 mL pipet and the green pipet pump.
VPCd. Into the 10-5, 10-6 and 10-7 sterile tubes, aseptically pipet exactly 9.0 mL sterile saline using a
10.0 mL pipet and the green pipet pump.
3. Make the first two tubes and the first plate transfers
a. Using a 1.0 mL pipet and the blue pipet pump, aseptically transfer exactly 0.1 mL of the original culture into the 10-2 dilution blank. Discard the pipet in the appropriate container. b. Roll the 10-2 tube between your hands to mix it. Then, with a fresh pipet, mix the culture further by drawing up some of the liquid at the bottom of the tube and releasing it at the top of the tube. Repeat the mixing several times. c. With the same pipet, aseptically transfer exactly 0.1 mL of the 10-2 dilution to the 10-4 blank.Discard the pipet in the appropriate container.
d. With a fresh pipet, mix the newly created 10-4 dilution tube by rolling it between your hands and by drawing the liquid up and down as described above. e. With the same pipet used for mixing, aseptically transfer exactly 1.0 mL of the 10-4 dilution into the 10-5 dilution blank. f. With the same pipet, transfer exactly 0.1 mL of the 10-4 dilution onto the 10-5 plate. Discard the pipet in the appropriate container. g. Spread the inoculum on the 10-5 plate according to the spread plate method.4. Make the next set of transfers from the 10-5 tube.
a. With a fresh pipet, mix the newly created 10-5 dilution tube by rolling it between your hands and by drawing the liquid up and down as described above. b. With the same pipet used for mixing, aseptically transfer exactly 1.0 mL of the 10-5 dilution to the 10-6 blank. c. With the same pipet, transfer exactly 0.1 mL of the 10-5 dilution onto the 10-6 plate. Discard the pipet in the appropriate container. d. Spread the inoculum on the 10-6 plate according to the spread plate method.5. Next set from the 10-6 tube.
a. With a fresh pipet, mix the newly created 10-6 dilution tube by rolling it between your hands and by drawing the liquid up and down as described above. b. With the same pipet used for mixing, aseptically transfer exactly 1.0 mL of the 10-6 dilution to the 10-7 blank. c. With the same pipet, transfer exactly 0.1 mL of the 10-6 dilution onto the 10-7 plate. Discard the pipet in the appropriate container. d. Spread the inoculum on the 10-7 plate according to the spread plate method.6. Next set from the 10-7 tube.
a. With a fresh pipet, mix the newly created 10-7 dilution tube by rolling it between your hands and by drawing the liquid up and down as described above. b. With the same pipet used for mixing, aseptically, transfer exactly 0.1 mL of the 10-7 dilution onto the 10-8 plate. Discard the pipet in the appropriate container. c. Spread the inoculum on the 10-8 plate according to the spread plate method.7. Allow the plates to stay right-side up for 5 minutes. Stack the plates in order and tape them
together with masking tape. Place them upside-down in the location designated for plates to be incubated. VPCProcedure Lab 2
1. Count the colonies on each plate and record the number in the data table. Colonies are best
counted by placing a dot on the bottom of the plate for each colony with a marker. If you get to100 colonies and have counted less than ¼ of the plate, and the growth is fairly even, you can
assume that there are more than 300 on the plate. Record ͞TNTC," too numerous to count in the colony column. If the area covered by 100 colonies is greater than ¼ of the plate, keep counting. If you reach 300 and still have more uncounted colonies, you can stop and record TNTC.2. Because each colony represents one CFU originally plated last week, you can determine the
number of CFUs. For plates with less than 30 colonies, you should record ͞TFTC," too few to count, in the CFU column. If the number of colonies was TNTC, then the number of CFUs is also TNTC.3. Calculate the Original Culture Density, OCD. If you have two countable plates, calculate an OCD
for each and then average the numbers. (Do not average the CFU numbers first.) Be sure to include the units in the answer. VPCLab Report: Viable Plate Count Part 1
Name ______________________________
Lab Section __________
Data and Observations
Plate Dilution
Factor
Number
of coloniesNumber of
CFUs represented Countable plate DF Countable plate CFU Original Cell Density (OCD)Second countable plate
if applicableOCD ____________________________
Post Lab Questions
1. Theoretically, how many countable plates should you have? Explain.
2. Give one case in which you could have two countable plates, theoretically.
VPC3. Note the results of a standard plate count below.
Plate Dilution
Factor
Number of
coloniesNumber of
CFUs represented10-4 1852
10-5 231
10-6 12
a. Calculate the OCD of the culture. Show your work. b. Explain the reasons that the 10-6 plate cannot be used in an accurate calculation of OCD. c. Explain why the 10-4 plate cannot be used in an accurate calculation of OCD.Post Lab Questions (Petersen, 2016)
1. You have a urine sample
from a patient that you suspect has a urinary tract infection. You make ten-fold dilutions of this sample as shown below, and then plate0.1 ml (100 µL) of the last
dilution on a TSA plate.There are 45 colonies on the
plate. How many CFUs/ml were in the original urine sample? VPC2. You have received a sample from a sewage treatment plant, and have been asked to determine
how many CFUs/ml are in this sample. You want to make a 1/100,000 fold dilution, but the smallest volume you can measure is 1.0 ml, and the tubes available to you only hold 10 ml.Draw a scheme showing how you would do this.
3. You do a series of dilutions as shown below, and you plate 1.0 ml of each dilution. Given the
information below, fill in the number of colonies you theoretically would expect on each of the plates. VPC4. You do serial dilutions on a water sample, and plate the dilutions on TSA plates. You count the
colonies on each of the plates as follows: (Note: TNTC = too numerous to count) PlateDilution
Factor
Number of
coloniesNumber of CFUs
represented10-2 Confluent
10-3 2044
10-4 352
10-5 50
10-6 6
10-7 0
Based on these results, what is your estimate for the total number of CFUs/ml in the original sample? Show your calculation. VPCLab Report: Viable Plate Count Part 2
Name ______________________________
Lab Section __________
Post Lab Questions
1. If your culture has 2.77 x 104 CFU/mL, what is the FDF that will yield a countable plate? (Hint:
Work backwards.) If you plate 0.1 mL, what tube dilution would it come from?2. If your culture has 6.5 x 107 CFU/mL, what FDF will yield a countable plate? If you plated 0.1 mL,
what tube dilution would it come from?3. A sample has between 9.41 x 105 and 9.41 x 108 CFU/mL. Devise a (one) dilution scheme (with
no extra plates) that will ensure you will get a countable plate. This should be a scheme doable with the supplies and measuring devices that we use in LSMCRB 121L. Make sure your scheme is legible! Be sure to show the following:Amounts in each dilution blank
Amounts transferred to each blank
Amounts transferred to each plate
Dilution factor of each dilution tube
Final dilution factor on each plate.
Count the minimum number of transfer pipettes by designating each pipette with a circled number and showing which transfers are made with each pipette as demonstrated in class.4. A sample has between 2.64 x 105 and 2.64 x 108 CFU/mL. Devise a (one) dilution scheme (with
no extra plates) that will ensure you will get a countable plate. This should be a scheme doable with the supplies and measuring devices that we use in LSMCRB 121L. Include all the information listed above. VPC5. A sample has between 1.27 x 106 and 6.86 x 109 CFU/mL Devise a (one) dilution scheme (with no
extra plates) that will ensure you will get a countable plate. This should be a scheme doable with the supplies and measuring devices that we use in LSMCRB 121L. Include all the information listed above.6. Given the parameters in question 5 above, devise a scheme that uses a minimum number of
dilution tubes and pipettes. VPCReferences
OpenStax CNX. (2018, Mar 19). OpenStax Microbiology. Retrieved from Petersen, J. a. (2016). Laboratory Excercises in Microbiology: Discovering the Unseen World Through Hands-On Investigation. CUNY Academic Works. Retrieved from http://academicworks.cuny.edu/qb_oers/16quotesdbs_dbs12.pdfusesText_18[PDF] how to calculate effective address in 8086
[PDF] how to calculate epinephrine dose
[PDF] how to calculate exchange rate
[PDF] how to calculate experimental yield
[PDF] how to calculate february days
[PDF] how to calculate floor area
[PDF] how to calculate freight cost
[PDF] how to calculate frequency spectrum
[PDF] how to calculate infusion time with drop factor
[PDF] how to calculate infusion time with tubing factor
[PDF] how to calculate logical address in 8086
[PDF] how to calculate mortgage payment on ba ii plus
[PDF] how to calculate multiple regression by hand
[PDF] how to calculate p value from chi square by hand
